Generate Data
group <- rep(c("treat", "placebo"),
each = 30)
symptom_post <- c(
rnorm(30, mean = 1, sd = 2),
rnorm(30, mean = 0, sd = 1)
)
dat1 <- data.frame(group, symptom_post)
Run Model
fit <- brm(bf(symptom_post ~ group, sigma ~ group),
data = dat1,
family = gaussian(),
seed = 1234) Family: gaussian
Links: mu = identity; sigma = log
Formula: symptom_post ~ group
sigma ~ group
Data: dat1 (Number of observations: 60)
Samples: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup samples = 4000
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 0.06698 0.15800 -0.24173 0.37999 1.00169 5280 2818
sigma_Intercept -0.17407 0.13197 -0.42005 0.10367 1.00107 3635 2690
grouptreat 0.71749 0.41627 -0.12423 1.53980 1.00066 2569 2354
sigma_grouptreat 0.90143 0.18605 0.53340 1.26824 1.00158 3215 2815
Samples were drawn using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).
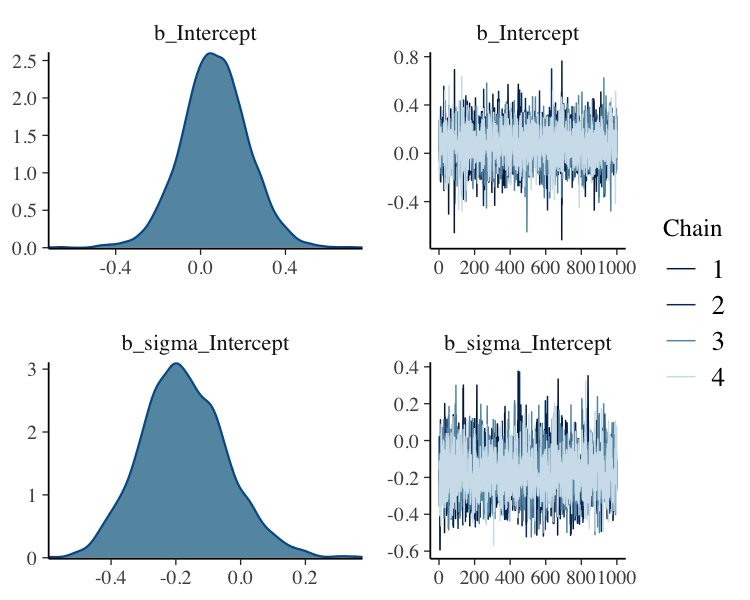
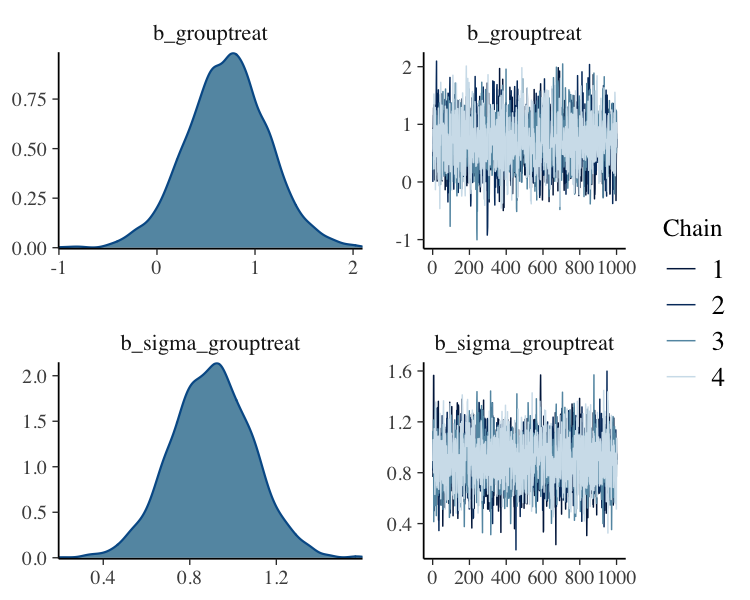
fit%>%
plot(N = 2, ask = FALSE)
Select first 100 Posterior Samples from each Chain
Family: gaussian
Links: mu = identity; sigma = log
Formula: symptom_post ~ group
sigma ~ group
Data: dat1 (Number of observations: 60)
Samples: 4 chains, each with iter = 1100; warmup = 1000; thin = 1;
total post-warmup samples = 400
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 0.07013 0.16137 -0.23785 0.38342 1.01057 774 364
sigma_Intercept -0.16386 0.14627 -0.42317 0.12234 1.00969 403 300
grouptreat 0.71854 0.41869 -0.04519 1.53400 1.01188 296 313
sigma_grouptreat 0.89130 0.19825 0.51367 1.30166 1.00422 344 263
Samples were drawn using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).
fit_slice%>%
plot(N = 2, ask = FALSE)
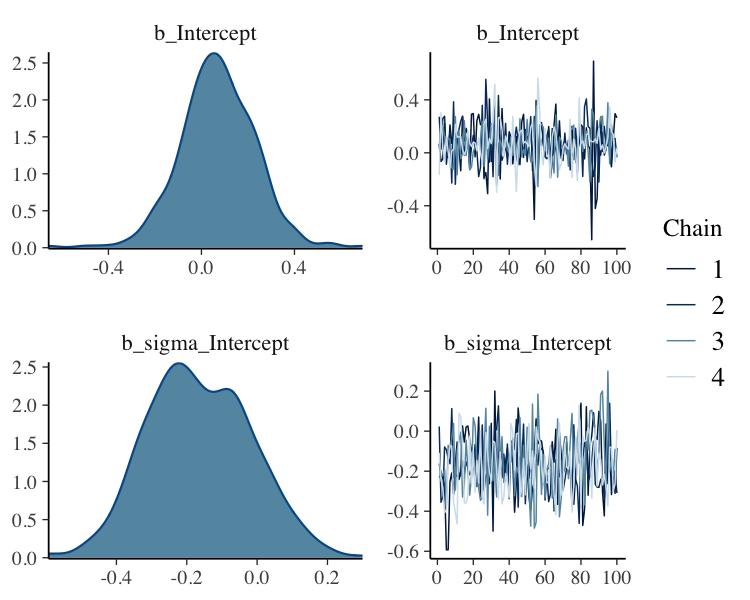
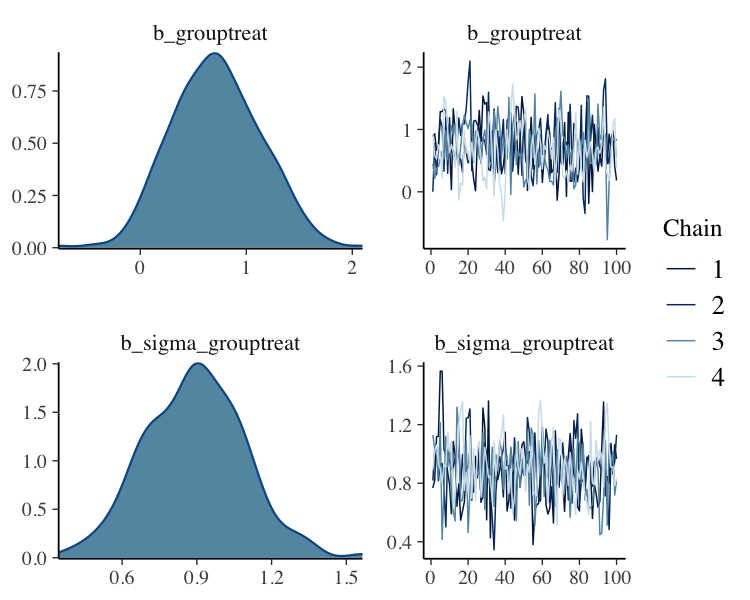
Thin 25% of Posterior Samples from each Chain
Family: gaussian
Links: mu = identity; sigma = log
Formula: symptom_post ~ group
sigma ~ group
Data: dat1 (Number of observations: 60)
Samples: 4 chains, each with iter = 1250; warmup = 1000; thin = 1;
total post-warmup samples = 1000
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 0.06683 0.15555 -0.24601 0.36279 1.00068 952 1033
sigma_Intercept -0.17761 0.13061 -0.42181 0.08630 1.00249 1022 1012
grouptreat 0.72207 0.41571 -0.07618 1.57548 1.00041 1003 894
sigma_grouptreat 0.90896 0.18475 0.56465 1.28181 1.00050 1052 994
Samples were drawn using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).
fit_thin%>%
plot(N = 2, ask = FALSE)
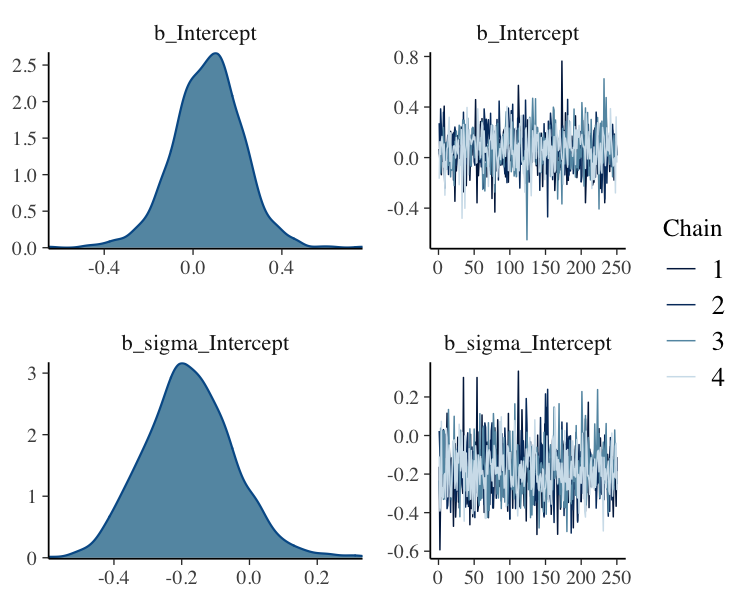
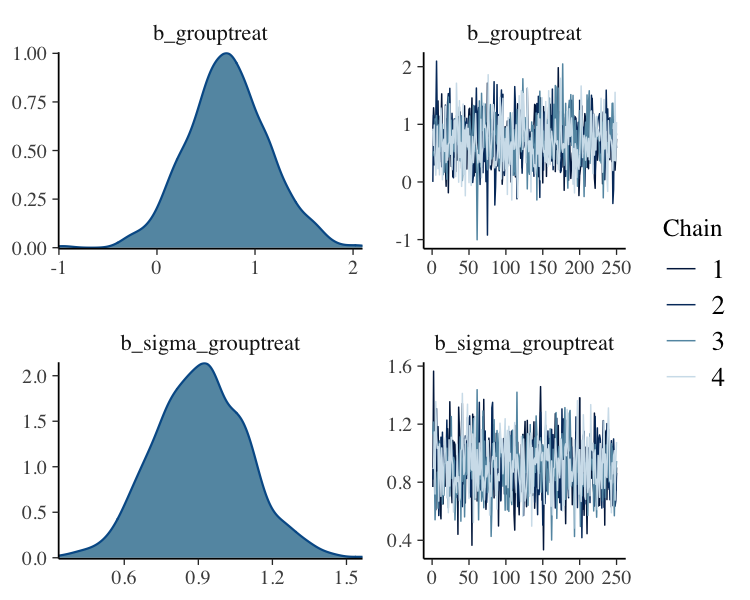
Filter Posterior Samples by Conditional
Family: gaussian
Links: mu = identity; sigma = log
Formula: symptom_post ~ group
sigma ~ group
Data: dat1 (Number of observations: 60)
Samples: 4 chains, each with iter = 1228; warmup = 1000; thin = 1;
total post-warmup samples = 912
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept -0.00769 0.15381 -0.32381 0.29970 1.00015 957 755
sigma_Intercept -0.17751 0.13422 -0.42817 0.13153 1.00122 659 794
grouptreat 1.25471 0.20926 1.01099 1.79655 1.00431 605 775
sigma_grouptreat 0.92356 0.20039 0.52929 1.30766 1.00501 614 655
Samples were drawn using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).
fit_filter%>%
plot(N = 2, ask = FALSE)
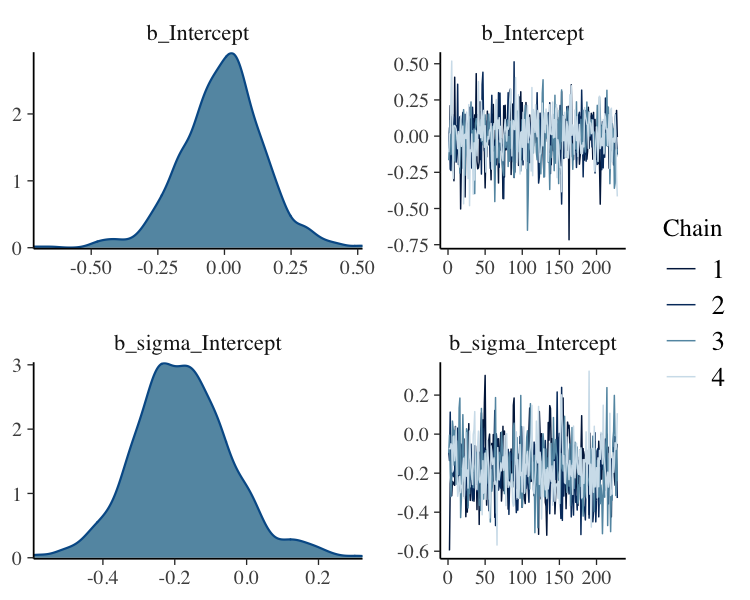
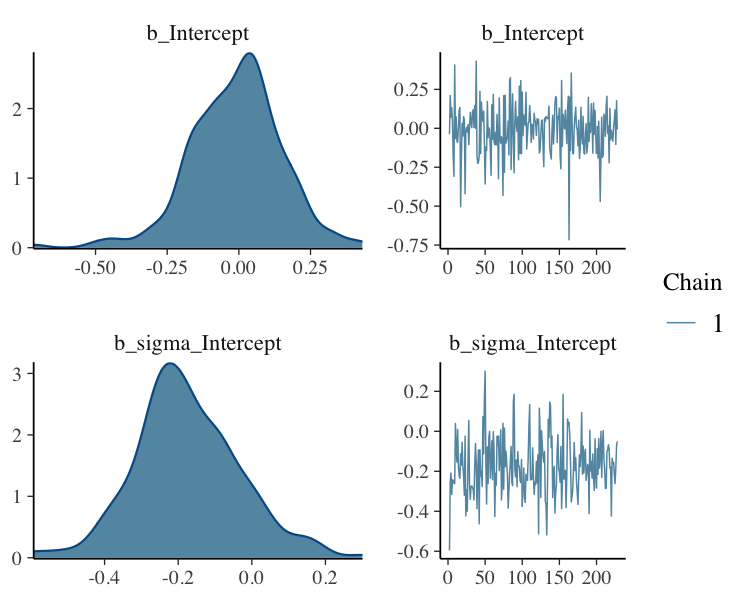
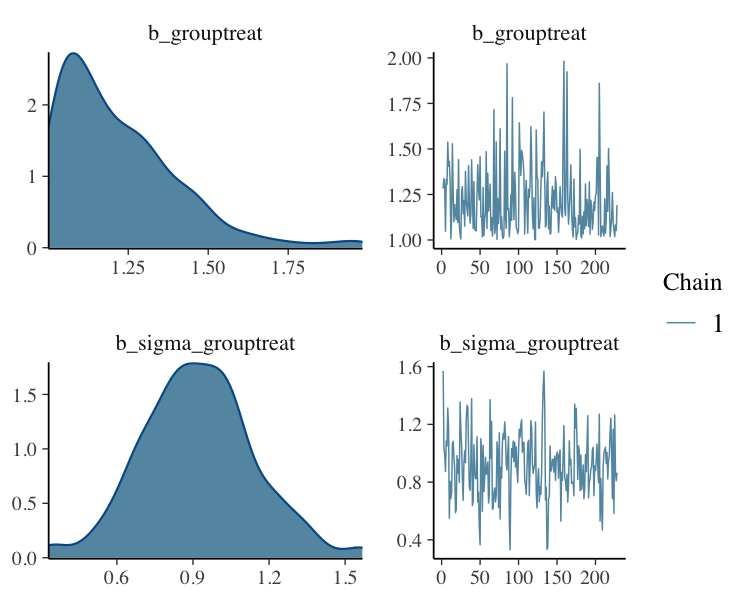
Filter Posterior Samples by Conditional Subset Chain
Family: gaussian
Links: mu = identity; sigma = log
Formula: symptom_post ~ group
sigma ~ group
Data: dat1 (Number of observations: 60)
Samples: 1 chains, each with iter = 1228; warmup = 1000; thin = 1;
total post-warmup samples = 228
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept -0.01025 0.16019 -0.33472 0.30657 1.00007 333 128
sigma_Intercept -0.17540 0.14239 -0.42458 0.13767 0.99844 157 153
grouptreat 1.21999 0.18977 1.00719 1.70702 0.99858 175 217
sigma_grouptreat 0.92037 0.22289 0.51294 1.35952 1.01292 139 140
Samples were drawn using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).
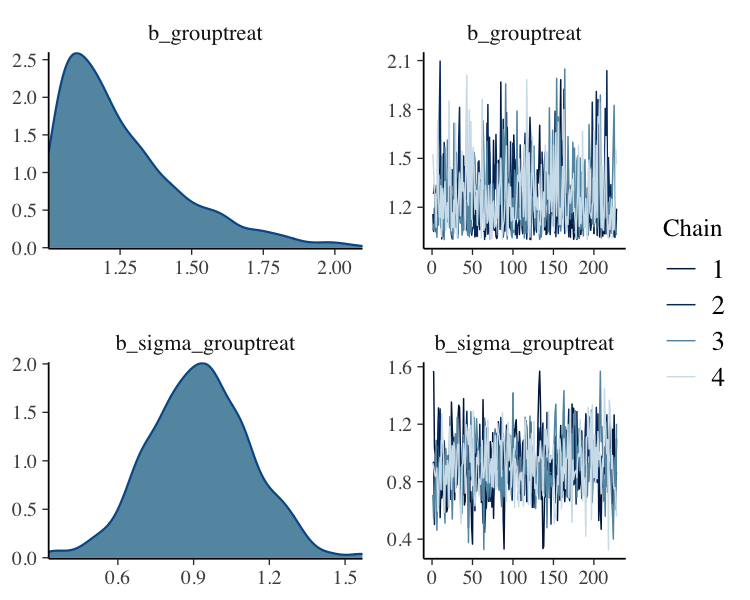
fit_filter_chain%>%
plot(N = 2, ask = FALSE)